Statistical Analysis by Two-Way ANOVA
You must select all (A) - (I) options to start a statistical analysis by two-way ANOVA.
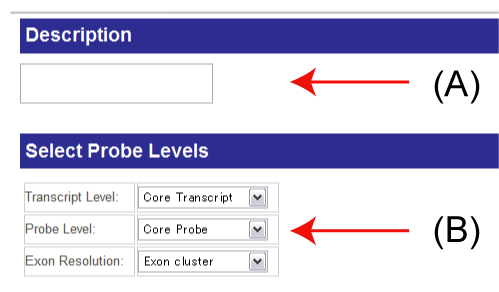
(A), (B)
(A): You can add a brief description of your analysis. It may be convenience that you put a name of this analysis to organize your analyses.
(B): You can select the level of expression information in exon array.
- Transcript level: For transcripts, there are three levels: core, extended and full transcripts according to their information quality.
- Probe level:
- Exon Resolution:
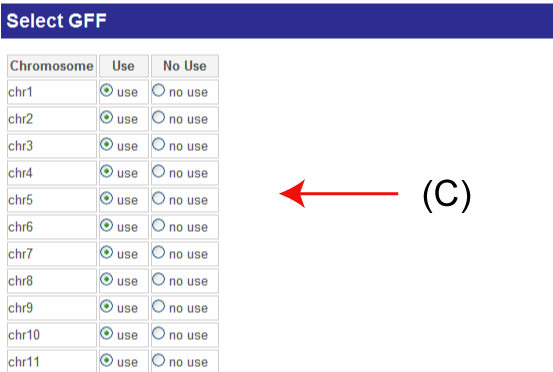
(C)
(C): You can select chromosomes. Transcripts on the selected chromosomes will be analyzed. This selection can reduce computational time.
(D), (E), (F)
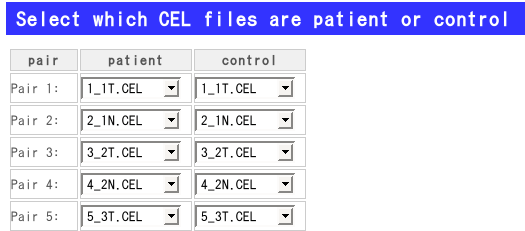
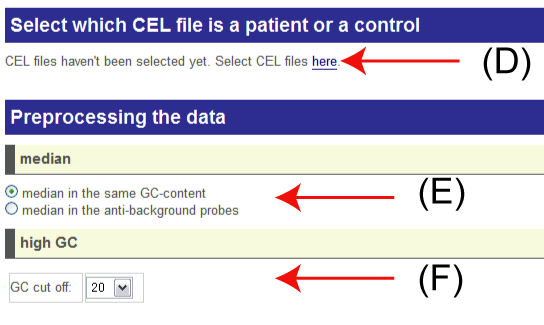
(D): Select the CEL files for the analysis that you have uploaded by FTP.
If you have already uploaded CEL files to analyze, you can see the following form. Then you can select pairs of CEL files.
(E): Select the type of normalization method.
- GC-content: uses median values in the same GC-content probe groups as control values.
- anti-background: uses median values in the same GC-content anti-background probes as control values.
(F): It is a possible case that probes with high GC-content work as noise. So you can remove such probes. In default, the probes with 20 or more GC-content are removed. If you want to use the all probes for analysis, yuou choose 25 as the cut-off.
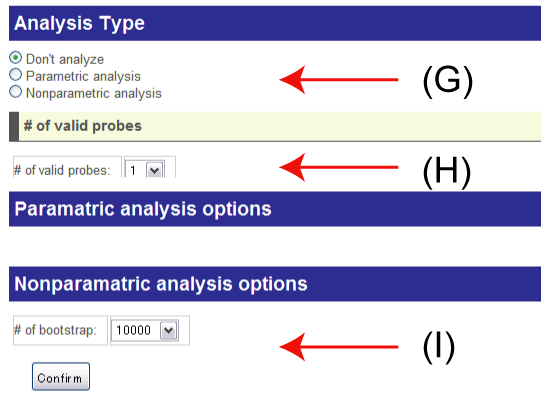
(G), (H), (I)
(G): Select the analysis types from the following three types:
- Don't analyze: ExonMiner does not perform ANOVA. Only visualization and sequence information are available.
- Parametric analysis: Gaussian distribution is assumed as the noise model.
- Nonparametric analysis: ExonMiner does not assume any distributions for the noise model. Bootstrap test will be applied for computing p-values.
(H): ExonMiner ignores probesets or exon clusters with small number of probes for stabilizing the results of ANOVA. You can choose this cut-off by this option.
(I): The number of bootstraps in nonparametric ANOVA.